Abstract
Allergic inflammation is based on the cross-linking of mast cell and basophil-bound IgE Abs and requires at least two binding sites for IgE on allergens, which are difficult to characterize because they are often conformational in nature. We studied the IgE recognition of birch pollen allergen Bet v 1, a major allergen for >100 million allergic patients. Monoclonal and polyclonal Abs raised against Bet v 1-derived peptides were used to compete with allergic patients’ IgE binding to Bet v 1 to search for sequences involved in IgE recognition. Strong inhibitions of patients’ IgE binding to Bet v 1 (52–75%) were obtained with mAbs specific for two peptides comprising aa 29–58 (P2) and aa 73–103 (P6) of Bet v 1. As determined by surface plasmon resonance, mAb2 specific for P2 and mAb12 specific for P6 showed high affinity, but only polyclonal rabbit anti-P2 and anti-P6 Abs or a combination of mAbs inhibited allergen-induced basophil degranulation. Thus, P2 and P6 define a surface patch on the Bet v 1 allergen, which allows simultaneous binding of several different IgE Abs required for efficient basophil and mast cell activation. This finding explains the high allergenic activity of the Bet v 1 allergen. The approach of using peptide-specific Abs for the mapping of conformational IgE epitopes on allergens may be generally applicable. It may allow discriminating highly allergenic from less allergenic allergen molecules and facilitate the rational design of active and passive allergen-specific immunotherapy strategies.
Type I allergy is an IgE-mediated hypersensitivity disease that affects >25% of the population worldwide (1). The simultaneous binding of IgE Abs to at least two different epitopes on allergens is a prerequisite for the cross-linking of mast cell and basophil-bound IgE Abs by allergens (2, 3). The cross-linking event initiates the release of inflammatory mediators, proteases, and proinflammatory cytokines from effector cells and causes immediate allergic inflammation and the well-known symptoms of allergic rhinoconjunctivitis, asthma, food allergy, skin inflammation, and life-threatening anaphylaxis (4). The sequences and three-dimensional structures of many important allergens have been determined in the last two decades and allowed analyzing IgE and T cell recognition of allergens (5, 6). IgE epitopes of major allergens seem to belong mainly to the conformational and/or discontinuous type that require the correct fold of the allergen for IgE binding and involve nonsequential amino acids (7, 8). Sequential IgE epitopes are rare and have been described mainly for food allergens, which may be due to the fact that true food allergens undergo proteolysis in the gastrointestinal tract before they induce an IgE sensitization. However, it has been shown that true food allergens also contain conformational epitopes (9–11). Several lines of evidence support the concept that IgE epitopes represent mainly conformational epitopes. First, it is almost impossible to map IgE epitopes of respiratory allergens using short synthetic allergen-derived peptides. Even large allergen fragments exhibit no or strongly reduced IgE reactivity compared with the intact folded allergen (12–14). Second, it has been shown that IgE recognition of allergens can be affected by alterations of the allergen conformation (e.g., calcium depletion from calcium-binding allergens, induction of structural changes through site-directed mutagenesis) (15–17). Two recently published structures of IgE allergen-immune complexes visualize the recognition of conformational epitopes by allergic patients IgE Abs (9, 18).
However, the mapping of conformational IgE epitopes of allergens remains a difficult task, and IgE epitopes of most of the important allergens, including the major allergen of birch, Bet v 1, have not yet been characterized in detail.
More than 100 million allergic patients, particularly in Northern and Middle Europe, North America, and Japan, are sensitized against the major birch pollen allergen, Bet v 1, a 17-kDa protein. Bet v 1-related allergens occur as cross-reactive allergens in trees belonging to the Fagales order (alder, hazel, hornbeam, oak) and in plant-derived food (e.g., apple, carrot) (19, 20). IgE recognition of Bet v 1 can be almost completely abolished when the protein is split into two unfolded fragments, each comprising approximately half of the Bet v 1 molecule (14). These results strongly suggest that IgE epitopes of Bet v 1 belong to the conformational type. Using site-directed mutagenesis, several authors have identified amino acids that may be involved in IgE recognition, but it is not clear whether these amino acids are directly involved in IgE recognition or contribute to subtle alterations in the allergen fold that cannot be visualized by methods such as circular dichroism, which study only the overall allergen fold (21–23). Furthermore, IgE recognition of the Bet v 1 mutants showed a considerable variability in individual patients. Information regarding the possible localization of IgE epitopes of Bet v 1 was obtained through Bet v 1-specific mAbs that inhibited IgE recognition of Bet v 1. An mAb that blocked IgE recognition to Bet v 1 reacted with a synthetic peptide comprising aa 49–66 of Bet v 1 (24). Furthermore, x-ray analysis of Bet v 1 in complex with a murine mAb that had yielded a moderate inhibition of IgE binding to Bet v 1 identified a conformational epitope containing adjacent aa E42, N43, I44, E45, G46, N47, G48, G49, P50, G51, and T52 and additional dispersed aa R70, D72, H76, I86, and K97 (22, 25).
In this study, we generated and identified mouse mAbs that were specific for two synthetic Bet v 1-derived peptides (peptide comprising aa 29–58 [P2] and aa 73–103 [P6]), which, after immunization of rabbits, had induced Abs that inhibited IgE binding to Bet v 1 almost as good as rabbit Abs raised against the complete Bet v 1 allergen (26). The mouse mAbs were used for the mapping of the epitopes recognized by allergic patients’ IgE using competitive binding assays. With this approach, a surface-exposed patch on the Bet v 1 structure was characterized that contained the majority of Bet v 1-specific IgE epitopes. Using peptide-specific monoclonal and polyclonal Abs for inhibition of Bet v 1-induced basophil degranulation, it is demonstrated that several different IgE Abs can bind simultaneously to the defined surface patch. Areas containing tightly clustered IgE-binding sites may be common features of potent allergens and represent determinants of allergenic activity.
Materials and Methods
Recombinant allergens, Bet v 1-derived peptides, rabbit Abs, and sera from allergic patients
Recombinant birch pollen allergen Bet v 1a, recombinant hypoallergenic isoform Bet v 1d, and Bet v 1-related recombinant allergens Aln g 1, Cor a 1, Mal d 1, Api g 1, and Dau c 1 were obtained from Biomay (Vienna, Austria) (27–33). Bet v 1-derived peptides (Table I) were synthesized and purified as described (26).
| Bet v 1-Derived Peptides . | Amino Acid Sequence . | Length (aa) . |
|---|---|---|
| P2 (aa 29–58) | LFPKVAPQAISSVENIEGNGGPGTIKKISFC | 30 |
| P2AI (aa 29–52) | LFPKVAPQAISSVENIEGNGGPGTI | 25 |
| P2BI (aa 35–58) | APQAISSVENIEGNGGPGTIKKISF | 25 |
| P2BII (aa 39–58) | SSVENIEGNGGPGTIKKISF | 20 |
| P2BIII (aa 44–58) | IEGNGGPGTIKKISF | 15 |
| P3 (aa 49–78) | CGPGTIKKISFPEGFPFKYVKDRVDEVDHTN | 30 |
| P6 (aa 73–103) | CEVDHTNFKYNYSVIEGGPIGDTLEKISNEIK | 31 |
| P6AI (aa 73–98) | EVDHTNFKYNYSVIEGGPIGDTLEKI | 26 |
| P6AII (aa 73–93) | EVDHTNFKYNYSVIEGGPIGD | 21 |
| P6AIII (aa 73–88) | EVDHTNFKYNYSVIEG | 16 |
| P6BI (aa 78–103) | NFKYNYSVIEGGPIGDTLEKISNEIK | 26 |
| P6BII (aa 83–103) | YSVIEGGPIGDTLEKISNEIK | 21 |
| P6BIII (aa 88–103) | GGPIGDTLEKISNEIK | 16 |
| Bet v 1-Derived Peptides . | Amino Acid Sequence . | Length (aa) . |
|---|---|---|
| P2 (aa 29–58) | LFPKVAPQAISSVENIEGNGGPGTIKKISFC | 30 |
| P2AI (aa 29–52) | LFPKVAPQAISSVENIEGNGGPGTI | 25 |
| P2BI (aa 35–58) | APQAISSVENIEGNGGPGTIKKISF | 25 |
| P2BII (aa 39–58) | SSVENIEGNGGPGTIKKISF | 20 |
| P2BIII (aa 44–58) | IEGNGGPGTIKKISF | 15 |
| P3 (aa 49–78) | CGPGTIKKISFPEGFPFKYVKDRVDEVDHTN | 30 |
| P6 (aa 73–103) | CEVDHTNFKYNYSVIEGGPIGDTLEKISNEIK | 31 |
| P6AI (aa 73–98) | EVDHTNFKYNYSVIEGGPIGDTLEKI | 26 |
| P6AII (aa 73–93) | EVDHTNFKYNYSVIEGGPIGD | 21 |
| P6AIII (aa 73–88) | EVDHTNFKYNYSVIEG | 16 |
| P6BI (aa 78–103) | NFKYNYSVIEGGPIGDTLEKISNEIK | 26 |
| P6BII (aa 83–103) | YSVIEGGPIGDTLEKISNEIK | 21 |
| P6BIII (aa 88–103) | GGPIGDTLEKISNEIK | 16 |
Cysteins added for coupling purposes are underlined.
Rabbit anti-Bet v 1, anti-P2, and anti-P6 Abs have been described and were affinity purified using protein G (Pierce, Rockford, IL) (26).
Sera from 44 patients who, according to case history, skin testing, and serology, were allergic to birch pollen were analyzed. The total IgE levels in the sera ranged from 24.1 to >5000 kUA/l and the birch pollen-specific IgE levels from 10.7 to >100 kUA/l according to CAP FEIA testing (Phadia, Uppsala, Sweden). Serum from a nonallergic person was included for control purposes.
Generation of peptide-specific mAbs
Synthetic Bet v 1-derived peptides P2 (aa 29–58) and P6 (aa 73–103) were coupled to keyhole limpet hemocyanin (KLH; Pierce) using a Conjugation Kit (Pierce). BALB/c mice (Charles River Laboratories, Sulzfeld, Germany) were immunized s.c. with the KLH-coupled Bet v 1-derived peptides (30 μg/ml per mouse), each adsorbed to Al(OH)3 (75 μl/mouse) (Serva, Heidelberg, Germany) on days 0, 28, and 46. Spleen cells of mice were harvested 3 d after final immunization. Hybridomas were raised by conventional hybridoma technology (34) with slight modifications, using the hypoxanthine, aminopterin, thymidine-sensitive, nonsecreting myeloma cell line X63Ag8.653 (35) as a fusion partner. Fused cells were suspended in hypoxanthine, aminopterin, thymidine medium (Sigma-Aldrich, Vienna, Austria), supplemented with feeder cells and initially seeded in 96-well tissue plates (Szabo, Vienna, Austria). Cells were allowed to grow for ∼2 wk and tested for the production of Bet v 1- and peptide-specific Abs by direct ELISA.
For this purpose, ELISA plates (Greiner, Kremsmünster, Austria) were coated with rBet v 1, P2 (aa 29–58), P6 (aa 73–103), KLH, and rPhl p 1 (5 μg/ml) diluted in PBS and blocked with 0.5% w/v BSA (Roth, Karlsruhe, Germany) in PBS + 0.05% Tween 20 for 1 h at 37°C. Hybridoma supernatants were added and incubated for 2 h at 37°C, and bound Abs were detected with a 1:1000 diluted rat anti-mouse IgG1 mAb (BD Pharmingen) for 2 h at 37°C followed by a 1:2000 diluted anti-rat IgG HRP-linked species-specific whole Ab (Amersham Biosciences, Uppsala, Sweden) for 30 min each at 37°C and 4°C. The plates were washed five times with PBS + 0.05% Tween 20 between incubation steps. Finally, plates were incubated with staining solution (ABTS diammonium salt; Sigma-Aldrich) at room temperature, and absorbance was measured at 405 nm on a microtiter plate reader (Dynatech, Denkendorf, Germany).
Confirmation of the IgG1 isotype of the mAbs was done using purified rat anti-mouse IgG1, IgG2a, IgG2b, IgM, and anti-IgE mAbs (BD Pharmingen).
Cross-reactivity of Bet v 1-specific mAbs was tested with rBet v 1a, rBet v 1d, rAln g 1, rCor a 1, rMal d 1, rApi g 1, and rDau c 1 (1 μg/ml) by ELISA in triplicate determinations, and results are displayed as mean OD values ± SD.
Inhibition of allergic patients’ IgE binding to allergens by Abs
The inhibition of allergic patients’ IgE binding to Bet v 1 by the mAbs was studied by ELISA inhibition experiments. ELISA plates were coated with rBet v 1 or related allergens (rAln g 1, rCor a 1, rMal d 1, rApi g 1) (1 μg/ml, diluted in PBS) at 4°C overnight. After blocking for 1 h at 37°C, plates were preincubated with culture supernatants from individual peptide-specific mAbs, mixes thereof, or with culture supernatant from an isotype-matched mAb (mAb4A6) specific for the unrelated allergen Bet v 2 (36). Then, plates were incubated with 1:5 diluted sera from birch pollen-allergic patients, and bound IgE Abs were detected with a 1:1000 diluted alkaline phosphatase-conjugated mouse monoclonal anti-human IgE Ab (BD Pharmingen) (37). The percentage inhibition of IgE binding to rBet v 1 after preincubation with Bet v 1-specific mAbs was calculated as follows: percent inhibition = 100 − (ODp × 100/ODnp). ODp and ODnp represent the extinctions after preincubation with hybridoma culture supernatant from Bet v 1-specific mAbs (ODp) and with the isotype control (ODnp), respectively.
Multiple sequence alignment
The sequences of Bet v 1a (Swissprot accession number: P15494), Bet v 1d (P43177), Aln g 1 (P38948), Cor a 1 (Q08407), Mal d 1 (Q9SYW3), Api g 1 (P49372), and Dau c 1 (O04298) were aligned using the multiple sequence alignment program Clustal W (38). The GeneDoc program (39) was used to visualize the degree of sequence identity among Bet v 1a-, Bet v 1d-, and Bet v 1-related allergens.
Basophil activation experiments
Whether Abs can inhibit allergen-induced activation of allergic patients’ basophils was investigated by measuring upregulation of CD203c activation (40). For this purpose, heparinized peripheral blood samples were obtained from birch-pollen allergic patients after informed consent was given. Blood samples were incubated (100 μl) for 15 min (37°C) with increasing concentrations (0.2–80 pM) of rBet v 1 that had been preincubated (4°C, overnight) with: 1) different concentrations of the individual Bet v 1-specific mAbs; 2) different concentrations of a mix of the two mAbs; or 3) different concentrations of a control mAb4A6. In addition, inhibition experiments were performed with different concentrations of Bet v 1-specific polyclonal Abs from Bet v 1-, P2-, and P6-immunized rabbits or IgG fractions from a rabbit immunized with a Bet v 1-unrelated Ag. For control purposes, cells were exposed to a monoclonal anti-IgE Ab (1 μg/ml; Immunotech, Marseille, France), buffer alone, or the Abs without addition of Bet v 1. The levels of CD203c expression were determined by flow cytometry and are expressed as stimulation index (SI) (means ± SDs of triplicate determinations) (40). Allergen-induced upregulation of CD203c was calculated from mean fluorescence intensities (MFIs) obtained with stimulated (MFIstim) and unstimulated (MFIcontrol) cells and expressed as SI (SI = MFIstim:MFIcontrol) (40).
The ability of Abs to inhibit allergen-induced basophil activation was also tested in another experimental system. Rat basophil leukemia cells (RBL-2H3 cells that had been transfected with the cDNA coding for the human high-affinity IgE receptor) (41) were sensitized with serum IgE from different birch pollen-allergic patients (42). RBL cell degranulation was then induced by the addition of increasing concentrations of rBet v 1 that had been preincubated (1 h, 37°C) either with: 1) Bet v 1-specific or isotype-matched mAbs; or 2) Bet v 1-specific polyclonal Abs or the corresponding preimmune IgG. The release of β-hexosaminidase from activated RBL cells was measured as described (3).
Competition ELISA with biotin-labeled mAbs
Immobilized Protein G [ImmunoPure (G) IgG Purification Kit; Pierce] was used to purify Bet v 1-specific monoclonal IgG1 Abs from hybridoma culture supernatants. The purified Abs were biotin-labeled using EZ-Link NHS-PEO4-Biotin (EZ-Link NHS-PEO4-Biotinylation Kit; Pierce) at a 20-fold molar excess of biotin for 30 min at room temperature. Buffer exchange and removal of free biotin was done with Desalt Spin Columns (Pierce).
ELISA plates were coated with 1 μg/ml rBet v 1 or P6 (aa 73–103) diluted in PBS (Maxisorp; Nunc) for 2 h at room temperature. After blocking, plate-bound allergen was either preincubated overnight at 4°C with increasing concentrations of unlabeled Bet v 1-specific mAbs (mAb2, 12, or 13) or, for control purposes, with an isotype-matched mAb (mAb4A6). After washing, plates were incubated with biotin-labeled Abs (mAb2-biotin, mAb12-biotin), and their binding was detected with streptavidin-peroxidase (Sigma-Aldrich) diluted 1:1000.
BIAcore analysis
The interaction of Bet v 1-specific mAbs (mAb2, mAb12) with rBet v 1 was measured by surface plasmon resonance using a BIAcore 2000 instrument (BIAcore, Uppsala, Sweden) at 25°C. The sensor chip C1 (BIAcore) surface was activated by injection of a 1:1 mixture of 1-ethyl-3-[3-dimethylaminopropyl]carbodiimide hydrochloride and N-hydroxysuccinimide at a flow rate of 5 μl/min for 7 min. Purified rBet v 1 (Biomay) was then covalently coupled to the activated surface by amine coupling. For this purpose, rBet v 1 was diluted in 10 mM sodium acetate (pH 4) (BIAcore) at a concentration of 200 μg/ml and was injected at 5 μl/min into flow cell 2. The unreacted active ester groups were blocked with 1 M ethanolamine (pH 8.5) at a flow rate of 5 μl/min for 7 min. The final immobilization level was 33.6 response units (flow cell 2). A blank surface (flow cell 1) containing no allergen was used as a reference. Binding experiments were carried out injecting purified Abs (c = 1 mg/ml) diluted in HBS-EP buffer (0.01 M HEPES, 0.15 M NaCl, 3 mM EDTA, 0.005% [v/v] surfactant P20 [pH 7.4]; BIAcore) at 2-fold increasing concentrations ranging from 0.52–16.5 nM. During the association phase, Bet v 1-specific mAbs were passed over the buffer-equilibrated chip surface (flow cells 1 and 2) at a rate of 30 μl/min for 5 min. Dissociation of Bet v 1-specific mAbs was investigated by injection of HBS-EP buffer at 30 μl/min for a period of 30 min. Regeneration of the sensor chip surface after each injection cycle was performed by a brief pulse (30 s, 20 μl/min) of 10 mM glycine-HCl (pH 2.5, mAb2; pH 2.25, mAb12).
Binding of the Bet v 1-specific mAbs (mAb2, mAb12) at different concentrations to the immobilized rBet v 1 was analyzed using BIAEvaluation 3.0 software (BIAcore). Association rate (ka), dissociation rate (kd), and affinity (KD) were calculated by numerical integration and fit to a 1:1 interaction model.
Epitope mapping with truncated peptides and building of a three-dimensional model
For epitope mapping, ELISA plates (Nunc) were coated with Bet v 1-derived peptides (P2, P2AI, P2BI, P2BII, P2BIII, P3, P6, P6AI, P6AII, P6AIII, P6BI, P6BII, and P6BIII) (Table I) and exposed to the Bet v 1-specific mAbs (mAb2, mAb4, mAb10, mAb12, and mAb13). Binding of Bet v 1-specific mAbs was detected with a 1:1000 diluted rat anti-mouse IgG1 (BD Pharmingen) followed by a 1:2000 diluted anti-rat IgG HRP-linked species-specific whole Ab (Amersham Biosciences). Results are displayed as mean ODs from triplicate determinations ± SD.
The PDB coordinates of the three-dimensional structure model of Bet v 1 [http://www.rcsb.org/pdb/explore/explore.do?structureId=1BV1 (43)] were used as a basis for the surface representation of Bet v 1 using the molecular graphics visualization tool RasMol (44). The minimal peptides reacting with mAb2, mAb12, and mAb13 were colored. For comparison, point mutations that yielded a reduction of IgE reactivity to Bet v 1 (21, 23) and binding sites of mAbs that had inhibited patients’ IgE binding to Bet v 1 (22, 24) have been also highlighted on the surface representation.
Results
Generation and characterization of mouse mAbs specific for peptides derived from the major birch pollen allergen Bet v 1
By immunization of BALB/c mice with Bet v 1-derived peptides P2 (aa 29–58) and P6 (aa 73–103) (Table I), which had been coupled to KLH, 14 hybridomas producing Bet v 1-specific mAbs were established and designated mAb1–14. The isotype and epitope specificity of the hybridomas was determined by direct ELISA using Bet v 1-derived peptides and isotype-specific Abs (Table II). Each of the 14 mAbs belonged to the IgG1 isotype and showed strong reactivity to the complete folded rBet v 1 molecule. mAbs raised against P2 (mAbs 1–11) reacted specifically with P2 but not with P6, and mAbs raised against P6 (mAbs 12–14) only recognized P6 but not P2. None of the mAbs reacted with the carrier protein KLH or the unrelated grass pollen allergen rPhl p 1.
| . | Isotype . | Epitope Specificity . | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| mAb No. . | IgG1 . | IgG2a . | IgG2b . | IgM . | IgE . | rBet v 1 . | P2 . | P6 . | rPhl p 1 . | KLH . |
| 1 | + | − | − | − | − | + | + | − | − | − |
| 2 | + | − | − | − | − | + | + | − | − | − |
| 3 | + | − | − | − | − | + | + | − | − | − |
| 4 | + | − | − | − | − | + | + | − | − | − |
| 5 | + | − | − | − | − | + | + | − | − | − |
| 6 | + | − | − | − | − | + | + | − | − | − |
| 7 | + | − | − | − | − | + | + | − | − | − |
| 8 | + | − | − | − | − | + | + | − | − | − |
| 9 | + | − | − | − | − | + | + | − | − | − |
| 10 | + | − | − | − | − | + | + | − | − | − |
| 11 | + | − | − | − | − | + | + | − | − | − |
| 12 | + | − | − | − | − | + | − | + | − | − |
| 13 | + | − | − | − | − | + | − | + | − | − |
| 14 | + | − | − | − | − | + | − | + | − | − |
| . | Isotype . | Epitope Specificity . | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| mAb No. . | IgG1 . | IgG2a . | IgG2b . | IgM . | IgE . | rBet v 1 . | P2 . | P6 . | rPhl p 1 . | KLH . |
| 1 | + | − | − | − | − | + | + | − | − | − |
| 2 | + | − | − | − | − | + | + | − | − | − |
| 3 | + | − | − | − | − | + | + | − | − | − |
| 4 | + | − | − | − | − | + | + | − | − | − |
| 5 | + | − | − | − | − | + | + | − | − | − |
| 6 | + | − | − | − | − | + | + | − | − | − |
| 7 | + | − | − | − | − | + | + | − | − | − |
| 8 | + | − | − | − | − | + | + | − | − | − |
| 9 | + | − | − | − | − | + | + | − | − | − |
| 10 | + | − | − | − | − | + | + | − | − | − |
| 11 | + | − | − | − | − | + | + | − | − | − |
| 12 | + | − | − | − | − | + | − | + | − | − |
| 13 | + | − | − | − | − | + | − | + | − | − |
| 14 | + | − | − | − | − | + | − | + | − | − |
Fourteen monoclonal Bet v 1-specific Abs were analyzed for their isotype and epitope-specificities by ELISA.
+, positive reaction; −, negative reaction.
Peptide-specific mAbs strongly inhibit binding of birch pollen-allergic patients’ IgE to the complete Bet v 1 allergen
To investigate if the peptide-specific mAbs can inhibit allergic patients’ IgE binding to the Bet v 1 allergen, ELISA competition experiments were performed (Table III). ELISA plate-bound Bet v 1 was preincubated with peptide-specific mAbs (mAb2, mAb4, mAb10, mAb12, or mAb13) and, for control purposes, with an isotype-matched mAb specific for an unrelated birch pollen allergen, Bet v 2. Table III shows the degree of inhibition of IgE binding achieved for sera from 34 birch pollen allergic patients. Each of the five Bet v 1-specific Abs caused, on average, a >50% inhibition of IgE binding. IgE binding to Bet v 1 was more strongly inhibited by Abs specific for P6 (mAb12 > mAb13) than by Abs specific for P2 (mAb2 > mAb4 > mAb10) (Table III).
| . | Inhibition (%) . | ||||
|---|---|---|---|---|---|
| Patient No. . | mAb2 . | mAb4 . | mAb10 . | mAb12 . | mAb13 . |
| 1 | 75 | 72 | 63 | 84 | 84 |
| 2 | 70 | 64 | 56 | 77 | 75 |
| 3 | 74 | 66 | 55 | 85 | 77 |
| 4 | 76 | 77 | 66 | 90 | 89 |
| 5 | 62 | 55 | 46 | 87 | 82 |
| 6 | 70 | 51 | 45 | 70 | 67 |
| 7 | 61 | 60 | 47 | 82 | 75 |
| 8 | 57 | 56 | 49 | 63 | 65 |
| 9 | 73 | 71 | 61 | 83 | 82 |
| 10 | 47 | 39 | 37 | 47 | 47 |
| 11 | 78 | 76 | 68 | 94 | 91 |
| 12 | 75 | 66 | 63 | 79 | 78 |
| 13 | 82 | 81 | 75 | 91 | 90 |
| 14 | 53 | 52 | 41 | 82 | 72 |
| 15 | 51 | 48 | 40 | 61 | 60 |
| 16 | 48 | 44 | 42 | 77 | 63 |
| 17 | 67 | 63 | 56 | 82 | 75 |
| 18 | 45 | 43 | 34 | 59 | 54 |
| 19 | 75 | 72 | 66 | 89 | 88 |
| 20 | 33 | 30 | 29 | 42 | 42 |
| 21 | 68 | 62 | 54 | 89 | 83 |
| 22 | 42 | 44 | 40 | 56 | 57 |
| 23 | 75 | 73 | 67 | 89 | 89 |
| 24 | 73 | 70 | 65 | 86 | 86 |
| 25 | 63 | 61 | 55 | 74 | 73 |
| 26 | 46 | 44 | 42 | 50 | 55 |
| 27 | 68 | 65 | 59 | 84 | 84 |
| 28 | 61 | 57 | 53 | 70 | 71 |
| 29 | 59 | 55 | 47 | 76 | 72 |
| 30 | 53 | 38 | 37 | 55 | 52 |
| 31 | 55 | 53 | 49 | 68 | 66 |
| 32 | 73 | 70 | 65 | 75 | 79 |
| 33 | 55 | 52 | 48 | 65 | 65 |
| 34 | 69 | 67 | 60 | 84 | 85 |
| Control | 4 | 0 | 0 | 0 | 0 |
| Mean | 63 | 59 | 52 | 75 | 73 |
| . | Inhibition (%) . | ||||
|---|---|---|---|---|---|
| Patient No. . | mAb2 . | mAb4 . | mAb10 . | mAb12 . | mAb13 . |
| 1 | 75 | 72 | 63 | 84 | 84 |
| 2 | 70 | 64 | 56 | 77 | 75 |
| 3 | 74 | 66 | 55 | 85 | 77 |
| 4 | 76 | 77 | 66 | 90 | 89 |
| 5 | 62 | 55 | 46 | 87 | 82 |
| 6 | 70 | 51 | 45 | 70 | 67 |
| 7 | 61 | 60 | 47 | 82 | 75 |
| 8 | 57 | 56 | 49 | 63 | 65 |
| 9 | 73 | 71 | 61 | 83 | 82 |
| 10 | 47 | 39 | 37 | 47 | 47 |
| 11 | 78 | 76 | 68 | 94 | 91 |
| 12 | 75 | 66 | 63 | 79 | 78 |
| 13 | 82 | 81 | 75 | 91 | 90 |
| 14 | 53 | 52 | 41 | 82 | 72 |
| 15 | 51 | 48 | 40 | 61 | 60 |
| 16 | 48 | 44 | 42 | 77 | 63 |
| 17 | 67 | 63 | 56 | 82 | 75 |
| 18 | 45 | 43 | 34 | 59 | 54 |
| 19 | 75 | 72 | 66 | 89 | 88 |
| 20 | 33 | 30 | 29 | 42 | 42 |
| 21 | 68 | 62 | 54 | 89 | 83 |
| 22 | 42 | 44 | 40 | 56 | 57 |
| 23 | 75 | 73 | 67 | 89 | 89 |
| 24 | 73 | 70 | 65 | 86 | 86 |
| 25 | 63 | 61 | 55 | 74 | 73 |
| 26 | 46 | 44 | 42 | 50 | 55 |
| 27 | 68 | 65 | 59 | 84 | 84 |
| 28 | 61 | 57 | 53 | 70 | 71 |
| 29 | 59 | 55 | 47 | 76 | 72 |
| 30 | 53 | 38 | 37 | 55 | 52 |
| 31 | 55 | 53 | 49 | 68 | 66 |
| 32 | 73 | 70 | 65 | 75 | 79 |
| 33 | 55 | 52 | 48 | 65 | 65 |
| 34 | 69 | 67 | 60 | 84 | 85 |
| Control | 4 | 0 | 0 | 0 | 0 |
| Mean | 63 | 59 | 52 | 75 | 73 |
The percentages of inhibition of IgE binding to rBet v 1 obtained with the Bet v 1-specific mAbs (mAb2, mAb4, mAb10, mAb12, mAb13) are displayed for 34 birch-pollen allergic patients and for serum from a nonallergic person. The mean percentages of inhibition are shown at the bottom of the table.
Interestingly, a combination of P2- and P6-specific mAbs did not lead to a strong increase of the inhibition of IgE binding compared with that obtained with P6-specific mAbs alone (data not shown).
mAb2 but not mAb12 shows extensive cross-reactivity with Bet v 1-homologous allergens
IgE Abs from Bet v 1-allergic patients show extensive cross-reactivity with Bet v 1-homologous allergens present in various tree pollen as well as in plant-derived food (19, 20). We therefore tested the reactivity of P2- and P6-specific mAbs with various Bet v 1-related proteins, among them cross-reactive allergens (Aln g 1, Cor a 1, Mal d 1, Api g 1, and Dau c 1) and a hypoallergenic Bet v 1 isoform (Bet v 1d) (Table IV). P2-specific mAb2 reacted with almost all tested proteins, whereas P6-specific mAb12 only reacted with Bet v 1 and the hypoallergenic isoform Bet v 1d (Table IV). Furthermore, we observed different reactivity profiles among the P2-specific mAbs. MAb 2 and mAb10 recognized Cor a 1 and Api g 1, whereas mAb4 did not (Table IV).
| . | mAb2 . | mAb4 . | mAb10 . | mAb12 . | mAb13 . | |||||
|---|---|---|---|---|---|---|---|---|---|---|
| Allergen . | OD . | ± SD . | OD . | ± SD . | OD . | ± SD . | OD . | ± SD . | OD . | ± SD . |
| Bet v 1 a | 1.118 | 0.101 | 1.083 | 0.034 | 0.986 | 0.026 | 1.141 | 0.005 | 1.157 | 0.014 |
| Bet v 1 d | 1.032 | 0.020 | 0.985 | 0.005 | 0.910 | 0.024 | 1.088 | 0.005 | 1.095 | 0.023 |
| Aln g 1 | 1.035 | 0.007 | 1.010 | 0.013 | 0.950 | 0.028 | 0.034 | 0.000 | 0.034 | 0.001 |
| Cor a 1 | 1.032 | 0.008 | 0.046 | 0.003 | 0.928 | 0.022 | 0.026 | 0.001 | 0.027 | 0.001 |
| Mal d 1 | 1.060 | 0.005 | 1.026 | 0.013 | 0.984 | 0.015 | 0.026 | 0.000 | 0.027 | 0.001 |
| Api g 1 | 0.912 | 0.016 | 0.024 | 0.001 | 0.779 | 0.018 | 0.023 | 0.001 | 0.022 | 0.000 |
| Dau c 1 | 0.027 | 0.001 | 0.027 | 0.001 | 0.027 | 0.001 | 0.024 | 0.001 | 0.026 | 0.003 |
| . | mAb2 . | mAb4 . | mAb10 . | mAb12 . | mAb13 . | |||||
|---|---|---|---|---|---|---|---|---|---|---|
| Allergen . | OD . | ± SD . | OD . | ± SD . | OD . | ± SD . | OD . | ± SD . | OD . | ± SD . |
| Bet v 1 a | 1.118 | 0.101 | 1.083 | 0.034 | 0.986 | 0.026 | 1.141 | 0.005 | 1.157 | 0.014 |
| Bet v 1 d | 1.032 | 0.020 | 0.985 | 0.005 | 0.910 | 0.024 | 1.088 | 0.005 | 1.095 | 0.023 |
| Aln g 1 | 1.035 | 0.007 | 1.010 | 0.013 | 0.950 | 0.028 | 0.034 | 0.000 | 0.034 | 0.001 |
| Cor a 1 | 1.032 | 0.008 | 0.046 | 0.003 | 0.928 | 0.022 | 0.026 | 0.001 | 0.027 | 0.001 |
| Mal d 1 | 1.060 | 0.005 | 1.026 | 0.013 | 0.984 | 0.015 | 0.026 | 0.000 | 0.027 | 0.001 |
| Api g 1 | 0.912 | 0.016 | 0.024 | 0.001 | 0.779 | 0.018 | 0.023 | 0.001 | 0.022 | 0.000 |
| Dau c 1 | 0.027 | 0.001 | 0.027 | 0.001 | 0.027 | 0.001 | 0.024 | 0.001 | 0.026 | 0.003 |
Positive reactions are bold.
Fig. 1 shows that the extensive cross-reactivity of mAb2 is associated with a high degree of sequence homology of peptide P2 to the homologous regions of the other allergens/proteins. P6 showed a considerably lower sequence homology. Overall, Aln g 1, Cor a 1, and Mal d 1 share >50% amino acid sequence identity with the major birch pollen allergen Bet v 1, whereas Api g 1 and Dau c 1 showed <50% amino acid identity.
Multiple sequence alignment of Bet v 1 isoforms (Bet v 1a, Bet v 1d) and Bet v 1-related allergens. Peptides comprising aa 29–58 (P2) and aa 73–103 (P6) of Bet v 1 are framed. Dots indicate identical amino acids, hyphens represent gaps, and asterisks mark every 10th amino acid. Percentages of sequence identity are indicated in brackets.
Multiple sequence alignment of Bet v 1 isoforms (Bet v 1a, Bet v 1d) and Bet v 1-related allergens. Peptides comprising aa 29–58 (P2) and aa 73–103 (P6) of Bet v 1 are framed. Dots indicate identical amino acids, hyphens represent gaps, and asterisks mark every 10th amino acid. Percentages of sequence identity are indicated in brackets.
Cross-reactive mAbs inhibit patients’ IgE binding to Bet v 1-related allergens
Major Bet v 1-related allergens from alder (Aln g 1), hazel (Cor a 1), apple (Mal d 1), and celery (Api g 1) were preincubated with mAb2, mAb4, or mAb10 prior to patients’ serum IgE. Each of the cross-reactive Bet v 1-specific mAbs inhibited allergic patients’ IgE binding to Aln g 1, Cor a 1, and Mal d 1 up to 75% (Table V). Only mAb4, which had not reacted with Cor a 1 and Api g 1, did not inhibit IgE reactivity to Cor a 1. No relevant inhibition of IgE binding to Api g 1 was observed with Bet v 1-specific mAb2, mAb4, and mAb10 (data not shown).
| . | Inhibition (%) rAln g 1 . | Inhibition (%) rCor a 1 . | Inhibition (%) rMal d 1 . | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Patient No. . | mAb2 . | mAb4 . | mAb10 . | mAb2 . | mAb4 . | mAb10 . | mAb2 . | mAb4 . | mAb10 . |
| 35 | 41 | 28 | 31 | 13 | 14 | 23 | 21 | 12 | 15 |
| 3 | 69 | 52 | 62 | 41 | 0 | 58 | 31 | 5 | 8 |
| 16 | 53 | 28 | 45 | 33 | 2 | 32 | 41 | 19 | 28 |
| 9 | 75 | 48 | 64 | 38 | 0 | 29 | 36 | 14 | 18 |
| 15 | 45 | 30 | 39 | 28 | 0 | 25 | 30 | 15 | 24 |
| 14 | 48 | 20 | 41 | 50 | 0 | 39 | 36 | 23 | 17 |
| 17 | 62 | 39 | 54 | 29 | 0 | 26 | 31 | 20 | 23 |
| 19 | 45 | 25 | 36 | 44 | 0 | 31 | 32 | 20 | 18 |
| Mean | 55 | 34 | 47 | 34 | 2 | 33 | 32 | 16 | 19 |
| . | Inhibition (%) rAln g 1 . | Inhibition (%) rCor a 1 . | Inhibition (%) rMal d 1 . | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Patient No. . | mAb2 . | mAb4 . | mAb10 . | mAb2 . | mAb4 . | mAb10 . | mAb2 . | mAb4 . | mAb10 . |
| 35 | 41 | 28 | 31 | 13 | 14 | 23 | 21 | 12 | 15 |
| 3 | 69 | 52 | 62 | 41 | 0 | 58 | 31 | 5 | 8 |
| 16 | 53 | 28 | 45 | 33 | 2 | 32 | 41 | 19 | 28 |
| 9 | 75 | 48 | 64 | 38 | 0 | 29 | 36 | 14 | 18 |
| 15 | 45 | 30 | 39 | 28 | 0 | 25 | 30 | 15 | 24 |
| 14 | 48 | 20 | 41 | 50 | 0 | 39 | 36 | 23 | 17 |
| 17 | 62 | 39 | 54 | 29 | 0 | 26 | 31 | 20 | 23 |
| 19 | 45 | 25 | 36 | 44 | 0 | 31 | 32 | 20 | 18 |
| Mean | 55 | 34 | 47 | 34 | 2 | 33 | 32 | 16 | 19 |
The percentages of inhibition of IgE binding to the Bet v 1-homologous allergens of alder (rAln g 1), hazel (rCor a 1) and apple (rMal d 1) by Bet v 1-specific mAbs (mAb2, mAb4, mAb10) are displayed for sera from eight birch-pollen allergic patients (numbered as in Table III). The mean percentages of inhibition are shown at the bottom of the table.
Bet v 1-induced basophil activation is inhibited by a mix of two peptide-specific monoclonal or by peptide-specific polyclonal IgG but not by the individual peptide-specific mAbs
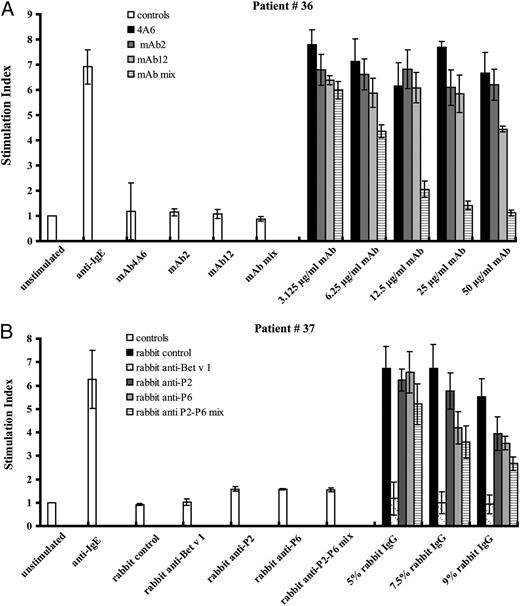
Next we investigated if the peptide-specific mAbs inhibit allergen-induced basophil activation. In a first set of experiments, basophils from three birch pollen-allergic patients were exposed to rBet v 1, which had been incubated with mAb2, mAb12, a mix of mAb2 and mAb12, or, for control purposes, with an isotype-matched control Ab (mAb4A6) (50 μg/ml), or medium alone (Fig. 2). Using the individual mAbs, no relevant inhibition of CD203c expression was observed in patients 36 and 38 and only a moderate inhibition in patient 37 at five different Bet v 1 concentrations (Fig. 2A–C). The mix of mAb2 and mAb12 almost completely suppressed Bet v 1-specific basophil activation in patient 36 and caused a moderate reduction in patient 37 (Fig. 2A–C). Similar results (i.e., no relevant inhibition of degranulation) were obtained when Bet v 1 was preincubated with the mAbs and the immune complexes were exposed to RBL cells expressing the human FcεRI (data not shown).
CD203c inhibition experiments. Blood samples from birch pollen-allergic patients were exposed to increasing concentrations of rBet v 1 (x-axes) that had been preincubated either with purified monoclonal Bet v 1-specific Abs (mAb2, mAb12, mix of mAb2 and mAb12) (A–C) or with purified polyclonal Bet v 1-specific Abs (rabbit anti-Bet v 1, rabbit anti-P2, rabbit anti-P6, rabbit anti-P2–P6 mix) or a purified rabbit control serum (D, E). CD203c expression on basophils was determined by FACS analysis and is displayed as SI on the y-axes.
CD203c inhibition experiments. Blood samples from birch pollen-allergic patients were exposed to increasing concentrations of rBet v 1 (x-axes) that had been preincubated either with purified monoclonal Bet v 1-specific Abs (mAb2, mAb12, mix of mAb2 and mAb12) (A–C) or with purified polyclonal Bet v 1-specific Abs (rabbit anti-Bet v 1, rabbit anti-P2, rabbit anti-P6, rabbit anti-P2–P6 mix) or a purified rabbit control serum (D, E). CD203c expression on basophils was determined by FACS analysis and is displayed as SI on the y-axes.
We therefore assumed that the lack of inhibition of IgE-mediated basophil activation by mAbs might be due to the fact that they do not compete with all of the IgE Abs that react with Bet v 1 in the areas defined by the peptides. Therefore, we tested whether polyclonal IgG Abs with specificity for the same peptides as those defined by the mAbs can inhibit allergen-induced activation of basophils. Indeed, we found that preincubation with anti-P2 and even more with anti-P6–specific polyclonal Abs resulted in a strong downregulation of CD203c expression induced by Bet v 1 in patients 36 and 38 (Fig. 2D, 2E). Interestingly, a mix of the polyclonal anti-P2 and anti-P6 Abs did not lead to a strong increase of the inhibition of CD203c upregulation. A very strong suppression of Bet v 1-induced CD203c upregulation was observed with polyclonal Abs raised against the complete Bet v 1 allergen (Fig. 2D, 2E).
In another set of experiments, we studied the influence of different concentrations of the monoclonal and polyclonal Abs on basophil activation induced by an optimal concentration of Bet v 1 (Fig. 3A, 3B). For patient 36, we found that already a mix of 12.5 μg/ml mAbs inhibited Bet v 1-induced basophil activation almost as well as the 50 μg/ml used for the inhibitions in Fig. 2, indicating that the lack of strong inhibition by the individual mAbs is not due to the use of too low Ab concentrations. Similar findings were made for the polyclonal peptide-specific Abs, confirming that the addition of 9% of the polyclonal peptide-specific Abs was sufficient to achieve inhibition of basophil activation.
CD203c inhibition experiments. Blood samples from birch pollen-allergic patients were exposed to rBet v 1 (x-axes) that had been preincubated either with different concentrations of purified monoclonal Bet v 1-specific Abs (mAb2, mAb12, mix of mAb2 and mAb12) (A) or with different concentrations of purified polyclonal Bet v 1-specific Abs (rabbit anti-Bet v 1, rabbit anti-P2, rabbit anti-P6, rabbit anti-P2–P6 mix) or a purified rabbit control serum (B). CD203c expression was determined by FACS analysis and is displayed in the form of SI on the y-axes.
CD203c inhibition experiments. Blood samples from birch pollen-allergic patients were exposed to rBet v 1 (x-axes) that had been preincubated either with different concentrations of purified monoclonal Bet v 1-specific Abs (mAb2, mAb12, mix of mAb2 and mAb12) (A) or with different concentrations of purified polyclonal Bet v 1-specific Abs (rabbit anti-Bet v 1, rabbit anti-P2, rabbit anti-P6, rabbit anti-P2–P6 mix) or a purified rabbit control serum (B). CD203c expression was determined by FACS analysis and is displayed in the form of SI on the y-axes.
In all experiments, the responsiveness of basophils to IgE-mediated stimulation was confirmed with anti-human IgE Abs, and incubation with buffer alone served as negative control. Furthermore, it is shown that none of the Abs caused direct activation of basophils without addition of allergen.
Simultaneous binding of peptide-specific mAbs to Bet v 1 and P6
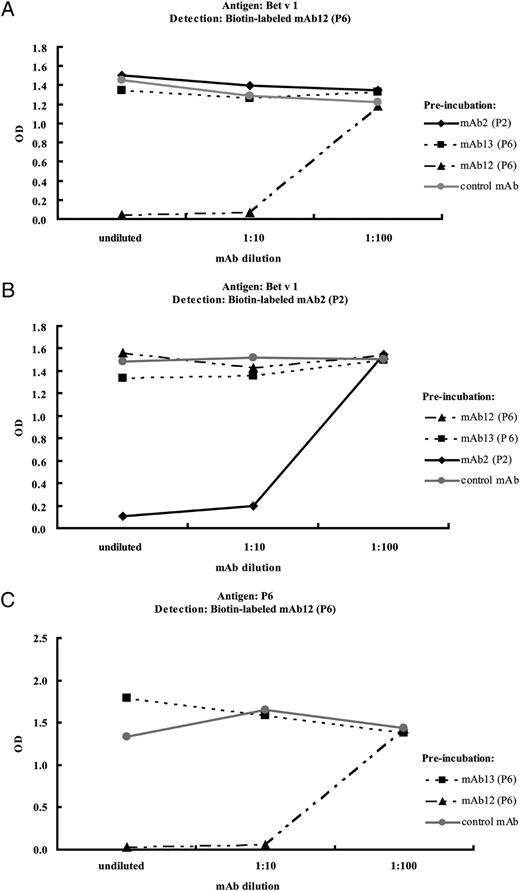
To test the hypothesis if indeed several Abs can bind simultaneously to the Bet v 1 allergen, we performed competitive ELISA assays. Using biotin-labeled mAb12 and mAb2, we investigated whether there is a simultaneous binding of the mAbs 2, 12, and 13 to Bet v 1. In a first set of experiments, Bet v 1 was preincubated with P6-specific mAb12, 13, P2-specific mAb2, and an isotype-matched control mAb and then exposed to the P6-specific biotin-labeled mAb12 (Fig. 4A) or with biotin-labeled P2-specific mAb2 (Fig. 4B). Our results demonstrate that neither preincubation with mAb2 nor with mAb13 reduced binding of biotin-labeled mAb12, although the peptide P2 is in close vicinity to peptide P6 on the Bet v 1 structure (Fig. 5), and mAb13 even reacts with the same 31 aa peptide as mAb12. mAb12 inhibited binding of biotin-labeled mAb12, demonstrating that the inhibition was performed at conditions of excess of competing Abs (Fig. 4A). A successful autoinhibition was also demonstrated for mAb2 (Fig. 4B). Preincubation of Bet v 1 with the isotype control did not influence the binding of biotin-labeled mAb2 or 12.
Simultaneous Ab binding to Bet v 1 and P6. Plate-bound rBet v 1 was preincubated with different dilutions of mAb2 (P2), mAb12 (P6), mAb13 (P6), or an isotype-control (mAb4A6) (x-axes) followed by incubation with biotin-labeled mAb12 (A) or mAb2 (B). In C, P6 was preincubated with different dilutions of mAb12 (P6) or mAb13 (P6) or the control Ab followed by incubation with biotin-labeled mAb12 (P6). The amounts of bound biotin-labeled mAbs correspond to the OD values (y-axes).
Simultaneous Ab binding to Bet v 1 and P6. Plate-bound rBet v 1 was preincubated with different dilutions of mAb2 (P2), mAb12 (P6), mAb13 (P6), or an isotype-control (mAb4A6) (x-axes) followed by incubation with biotin-labeled mAb12 (A) or mAb2 (B). In C, P6 was preincubated with different dilutions of mAb12 (P6) or mAb13 (P6) or the control Ab followed by incubation with biotin-labeled mAb12 (P6). The amounts of bound biotin-labeled mAbs correspond to the OD values (y-axes).
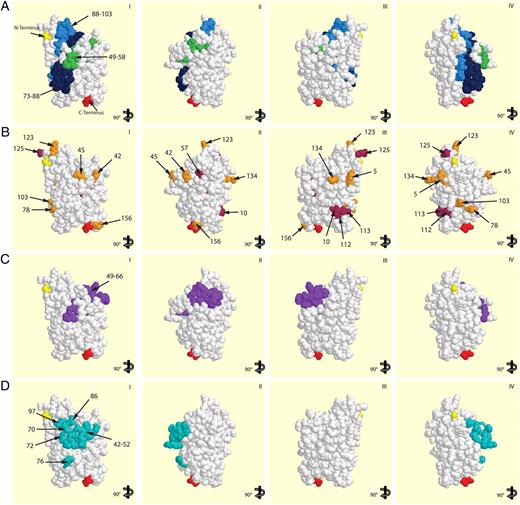
Localization of amino acid residues on the Bet v 1-structure involved in allergic patient’ IgE recognition. A shows a surface representation of the Bet v 1 molecule with the minimal peptides binding mAb2 (aa 49–58, green), mAb12 (aa 88–103, light blue), and mAb13 (aa 73–88, dark blue). B shows the localization of aa 10, 30, 57, 112, 113, 125 (maroon) and aa 5, 42, 45, 78, 103, 123, 134, 156 (orange), which, according to in vitro mutagenesis studies, were supposed to be involved in IgE recognition. C illustrates aa 49–66, and D shows aa 42–52, 70, 72, 76, 86, and 97, which are involved in binding of mAbs that inhibited IgE binding to Bet v 1. I–IV, Different views of the Bet v 1 surface. N (yellow) and C termini (red) of Bet v 1 are indicated.
Localization of amino acid residues on the Bet v 1-structure involved in allergic patient’ IgE recognition. A shows a surface representation of the Bet v 1 molecule with the minimal peptides binding mAb2 (aa 49–58, green), mAb12 (aa 88–103, light blue), and mAb13 (aa 73–88, dark blue). B shows the localization of aa 10, 30, 57, 112, 113, 125 (maroon) and aa 5, 42, 45, 78, 103, 123, 134, 156 (orange), which, according to in vitro mutagenesis studies, were supposed to be involved in IgE recognition. C illustrates aa 49–66, and D shows aa 42–52, 70, 72, 76, 86, and 97, which are involved in binding of mAbs that inhibited IgE binding to Bet v 1. I–IV, Different views of the Bet v 1 surface. N (yellow) and C termini (red) of Bet v 1 are indicated.
Next we investigated if mAb12 and mAb13 can bind simultaneously to the same 31 aa P6 peptide (Fig. 4C). We found that mAb12 but not mAb13 inhibited binding of biotin-labeled mAb12 to the P6 peptide, confirming that both mAbs bind to P6 simultaneously.
High-affinity binding of peptide-specific mAbs to Bet v 1
Another possibility for the lack of inhibition of Bet v 1-induced basophil activation by the peptide-specific mAbs might be that they exhibit only a low affinity for Bet v 1. We therefore measured the affinity constants of P2-specific mAb2 and P6-specific mAb12 to the major birch pollen allergen Bet v 1 using surface plasmon resonance measurements (Fig. 6). Two-fold increasing concentrations (mAb2: 0.52–4.13 nM; mAb12: 2.06–16.5 nM) of purified Bet v 1-specific mAbs were injected into the flow cell containing immobilized rBet v 1, and binding as well as dissociation were recorded in real-time. Fig. 6 shows the binding sensograms with an overlay of recorded (colored) and calculated (black) curves. ka and kd were determined to be ka = 2.39e6/Ms and kd = 2.00e-4/s for mAb2 and ka = 2.09e5/Ms and kd = 2.18e-4/s for mAb12. The dissociation constant KD was calculated from corresponding ka and kd to be 8.35e-11 M for mAb2 and 1.05e-9 M for mAb12. The affinities of both mAbs are therefore comparable to those recorded for human allergen-specific IgE Abs (9, 18, 45).
Affinity and kinetic analysis of the interaction between rBet v 1 and the Bet v 1-specific mAbs mAb2 (P2) and mAb12 (P6) using surface plasmon resonance (Biacore) measurements. mAb2 (A) and mAb12 (B) were injected at 2-fold increasing concentrations into the flow cell containing immobilized rBet v 1 (first arrow). The end of injection is indicated by a second arrow, followed by monitoring of Ab dissociation. The binding curves (colored) were fitted (black curves) according to a 1:1 interaction model. Binding in A and B is expressed in response units (RU) on the y-axes over time.
Affinity and kinetic analysis of the interaction between rBet v 1 and the Bet v 1-specific mAbs mAb2 (P2) and mAb12 (P6) using surface plasmon resonance (Biacore) measurements. mAb2 (A) and mAb12 (B) were injected at 2-fold increasing concentrations into the flow cell containing immobilized rBet v 1 (first arrow). The end of injection is indicated by a second arrow, followed by monitoring of Ab dissociation. The binding curves (colored) were fitted (black curves) according to a 1:1 interaction model. Binding in A and B is expressed in response units (RU) on the y-axes over time.
Detailed epitope mapping of the peptide-specific mAbs
Next we performed a detailed epitope mapping of the Bet v 1-specific mAbs (mAb2, mAb4, mAb10, mAb12, and mAb13) using a series of truncated peptides (Table VI). A truncation of P2 for 5 amino acids at the N- or C-terminal end indicated that the 5 C-terminal amino acids (KKISF) are critical for binding of mAb2, 4, and 10 (Table VI). Finally, a 10-mer peptide (aa 49–58; indicated in green) was identified as minimal peptide capable of binding mAbs 2, 4, and 10 (Table VI).
 |
 |
Minimal binding sequence for peptide 2-specific mAbs (green) and peptide 6-specific mAbs (blue). Positive reactions are bold.
The P6-specific mAbs (mAb12, mAb13) recognize different fragments within P6 (aa 73–103). mAb12 bound to a P6-derived C-terminal 16-mer peptide comprising aa 88–103 (indicated in light blue), whereas mAb13 reacted with a P6-derived N-terminal 16-mer comprising aa 73–88 (indicated in dark blue) of Bet v 1 (Table VI).
Localization of the minimal binding sites of mAbs 2, 12, and 13 on the Bet v 1 molecule
Fig. 5A shows a surface representation of the Bet v 1 molecule with the proteins’ N and C terminus colored in yellow and red, respectively. The minimal peptides binding mAb2 (green), mAb12 (light blue) and mAb13 (dark blue) are located in close vicinity on the Bet v 1 surface. According to the binding experiments with the biotin-labeled Abs and the inhibition of basophil activation with polyclonal anti-peptide Abs, at least three, if not more, different complete Abs can bind simultaneously to the area defined by the peptides. Amino acids that, according to earlier in vitro mutagenesis studies, are involved in IgE recognition of Bet v 1 have been highlighted on the Bet v 1 structure in Fig. 5B (21, 23). Serine 57, asparagine 78, and lysine 103 that, according to mutagenesis studies, are involved in IgE recognition of Bet v 1 are part of P2 and P6, respectively. However, most of the other amino acids that, according to mutagenesis studies, affect IgE recognition are spread over different sides of the Bet v 1 allergen, and not all of them (e.g., phenylalanine 30) are surface-exposed.
Fig. 5C shows peptide 49–66 representing the binding site of mAbs that had inhibited patients’ IgE binding to Bet v 1 (24). In Fig. 5D, residues involved in the binding of another mAb that reportedly had caused a 40% inhibition of IgE binding to Bet v 1 are highlighted (22, 25). The peptides and amino acid residues defined in the latter studies performed with other mAbs are in close vicinity and/or overlap with the area defined by P2 and P6.
Discussion
Whether a protein Ag (i.e., allergen) induces strong immediate type inflammatory reactions in sensitized allergic patients is primarily associated with its ability to induce IgE-mediated degranulation of mast cells and basophils (46). The degranulation event is based on the cross-linking of cell-bound IgE Abs and hence requires the presence of at least two IgE epitopes on the allergen (2, 3). The sequences and three-dimensional structures of many important allergens have been elucidated in the last 20 years, but our knowledge regarding the precise mode of how IgE Abs recognize allergens is still limited due to the fact that IgE Abs seem to recognize primarily conformational epitopes on allergens (8). Conformational epitopes cannot be easily mapped using conventional epitope-mapping strategies such as testing for IgE reactivity to recombinant or synthetic allergen fragments because fragmentations of proteins often leads to the loss of the three-dimensional structure of the protein and hence to loss of IgE reactivity (14, 47). The technology of in vitro mutagenesis may highlight amino acids that are involved in IgE binding, but it is difficult to determine whether a certain mutation affects IgE Ab reactivity because it affects the allergen’s conformation or because it affects directly the interaction with the V region of IgE (21, 48).
In two recent studies, the three-dimensional structure of an allergen in complex with a corresponding human IgE Ab has been reported (9, 18). These studies have indeed visualized the conformational recognition of allergens by IgE, but the technique of cocrystallization does not allow analyzing the repertoire of a polyclonal IgE response toward an allergen.
In this study, we have used peptide-specific monoclonal and polyclonal Abs for an indirect mapping of the IgE binding sites of one of the most potent respiratory allergens, the major birch pollen allergen Bet v 1. This technology has been described for the mapping of conformational epitopes in certain experimental model systems, but as yet has not been fully explored for the mapping of conformational IgE epitopes on allergens (7). We generated mAbs specific for two Bet v 1 peptides, P2 (aa 29–58) and P6 (aa 73–103), which are not adjacent in the sequence of Bet v 1 but appear in close vicinity on the three-dimensional structure of Bet v 1. mAbs specific for these peptides inhibited strongly (>50%) the binding of allergic patients’ IgE Abs to Bet v 1. This finding is interesting for two reasons. First, it confirms that single mAbs can strongly inhibit the polyclonal IgE recognition of allergic patients. Second, neither P2 nor P6 themselves show any reactivity with allergic patients’ IgE. This may be explained by the fact that the peptides are either only parts of IgE epitopes or lie in close vicinity to the IgE epitopes so that the corresponding mAbs can cause an inhibition of IgE binding by steric hindrance. Using truncated peptides, the binding sites for the mAbs could be mapped to sequences aa 49–58, aa 73–88, and aa 88–103, which defined a patch on the Bet v 1 allergen. This patch includes several amino acids that, in previous in vitro mutagenesis studies, have been identified as important amino acids involved in IgE binding (21, 23). Furthermore, the area defined in our study overlaps with the binding sites of other mAbs that had inhibited patients’ IgE binding to the Bet v 1 allergen (22, 24, 25). Two earlier studies describe the complex of a Bet v 1-specific Fab with the Bet v 1 allergen. The Fab was derived from a mouse monoclonal anti-Bet v 1-specific Ab (BV16), which, unlike our Abs, recognized a conformational epitope on Bet v 1-containing adjacent aa E42, N43, I44, E45, G46, N47, G48, G49, P50, G51, and T52 and additional dispersed aa R70, D72, H76, I86, and K97 (22, 25).
Lebecque et al. (24) described a panel of mouse monoclonal anti-Bet v 1 Abs obtained by immunizing mice with birch pollen grains. Four of these Abs inhibited allergic patients’ IgE binding to Bet v 1 strongly and recognized a peptide epitope comprising G49–Y66. The agreement of our results with the earlier studies indicates that the patch characterized by us indeed contains the major binding sites for Bet v 1-specific IgE Abs. We are also quite confident that the patch on the Bet v 1 molecule defines a major IgE epitope-containing area because we could inhibit a high percentage (>50%) of the binding of Bet v 1-specific IgE Abs from a large number of allergic patients with the P2 and P6-specific mAbs.
However, much to our surprise, we found that neither mAb2 nor mAb12 alone caused any relevant inhibition of Bet v 1-induced basophil activation, although IgE binding to the allergen was strongly reduced. This finding was surprising because several other allergen-specific mAbs that had inhibited IgE reactivity to the corresponding allergen [e.g., grass pollen allergen Phl p 2; birch pollen allergen, Bet v 1 (18, 37, 49)] to a similar extent as the mAbs described by us had also inhibited allergen-induced basophil degranulation.
As one possible explanation for the lack of inhibition of basophil activation by the mAbs, we initially considered low affinity of the peptide-specific Abs to the folded Bet v 1 allergen. However, plasmon surface resonance experiments showed that mAb2 and mAb12 bound to Bet v 1 with good affinity, which was almost as high as that recorded for human allergen-specific IgE Abs (9, 18, 45).
When we then compared the ability of the monoclonal peptide-specific Abs with polyclonal Abs raised against the very same peptides, we found that the peptide-specific polyclonal Abs inhibited quite well allergen-induced basophil activation. A partial inhibition was also obtained with a mix of mAb2 and mAb12. These findings indicated that the patch defined by the mAbs contains binding sites for several different IgE Abs for which binding is not efficiently enough blocked by one or two mAbs, whereas polyclonal peptide-specific IgG Abs populate the area more densely and thus cause efficient inhibition. In this context, it should be noted that when Bet v 1 was captured by mAb2 or mAb12, polyclonal anti-P2 as well as anti-P6 Abs still bound to Bet v 1, indicating that the polyclonal antisera contained Abs that could bind in addition to the mAbs to Bet v 1 (data not shown). Because the size of IgE and IgG Abs is ∼10-fold that of the Bet v 1 allergen, it seemed quite surprising that several different Abs may be able to bind to Bet v 1 and in particular to the relatively small patch defined by mAb2 and mAb12 on the Bet v 1 structure. However, in competitive ELISA experiments, we could indeed show that two different IgG Abs can bind simultaneously to the IgE-reactive patch defined by mAb2 and mAb12.
Our results thus indicate that several (i.e., two or more) IgE-binding sites cluster within a relatively small area on the Bet v 1 allergen. This finding may explain why Bet v 1 is a highly potent allergen that induces strong degranulation of basophils and mast cells and thus immediate allergic inflammation in patients.
To the best of our knowledge, our study is the first to use peptide-specific mAbs to map conformational IgE epitopes on an important allergen and to suggest this strategy as a generally applicable technology for the long-sought goal of mapping conformational IgE epitopes on allergens. Another important, novel, and unexpected finding of our study is the demonstration that several different IgE Abs can bind simultaneously to a small surface patch on the Bet v 1 allergen. This finding is surprising considering the size of a complete IgE molecule (>180 kDa) versus the size of a patch comprising only the small portion of the 17-kDa allergen molecule defined by peptides P2 and P6.
The knowledge of the localization of IgE epitopes on the Bet v 1 allergen should also contribute to the design of effective active and passive immunotherapy strategies for birch pollen and related allergies. Finally and perhaps most important, the technology of using peptide-specific Abs for indirect IgE epitope mapping may be generally applicable and allow elucidating the IgE epitopes of many important other allergens for the design of allergen-specific therapy strategies.
Footnotes
This work was supported by the Christian Doppler Research Association, Biomay and by Grants F1809 and F1815 from the Austrian Science Fund (Fonds zur Förderung der Wissenschaftlichen Forschung).
References
Disclosures
The authors have no financial conflicts of interest.